BLIPKIT
A usage strategy emerges
Thu, 2009-11-12 17:31 | by Samuel LampaA strategy for how to work with the Bioclipse/JPL/Prolog/Blipkit combination I'm setting up, is becoming clear.
The main idea with Bioclipse, as well as with having a prolog engine available in it, is for flexible and "interactive" knitting together of knowledge. One of the main questions regarding how to use a Bioclipse/JPL/Prolog/Blipkit combination, has been where to put the bulk of knowledge integration/reasoning code? There would in principle be three options for that:
- Bioclipse (Javascript environment)
- The Blipkit-Prolog/Bioclipse integration plugin (Java code, a.k.a. "Manager methods")
- The prolog engine (As a prolog file)
How to deal with rdf namespaces (does not work well with JPL)
Mon, 2009-11-09 18:09 | by Samuel LampaI had the problem that in JPL (The java Prolog API) you cannot use namespaces before term (atoms or variables etc.) names, like so:
prologFunction( ns:'atom' ).
The best solution would be to have some kind of "namespace-like" support in the JS console of Bioclipse instead. One easy thing one can do is to just create a simple function that appends the long preceding URL, so a JS Example could be:
function molid ( term ) {
return "http://pele.farmbio.uu.se/nmrshiftdb/?moleculeId=" + term;
}
blipkit.queryRDF(molid("234"),"X","Y");Pondering over strategies for evaluating BLIPKIT vs Pellet for use with NMRShift data
Tue, 2009-11-03 10:48 | by Samuel LampaAs a subproject, I'm now focusing on doing some comparison between reasoning in Prolog and Pellet, using example data from NMRShiftDB.org. Below I'm documenting my planning, + any new findings while I'm digging in to this subpeoject.
Testing out querying Prolog from Bioclipse
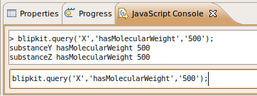
Sun, 2009-11-01 00:20 | by Samuel LampaMy project has been more or less on hold for around a week because of exams and other stuff. Looking forward to getting some concrete things done now. While still reading up a bit on OWL, I've started taken the first steps of the prolog/blipkit integration with a simple method for querying a prolog on the form "subject, a predicate, and an object".
Accessing BLIPKIT from command line Java or Eclipse Plug-in in Ubuntu
Fri, 2009-10-16 20:28 | by Samuel LampaFor my own documentation I went ahead and summarized all the steps I had to take
- In order to start blipkit from java commandline, and
- In order to start blipkit from an Eclipse plugin.
Got blipkit started from inside Eclipse
Fri, 2009-10-16 14:55 | by Samuel LampaI now also managed to start blipkit from inside Eclipse.
The trick was to start the whole eclipse (The eclipse using for building Bioclipse) preceded with LD_PRELOAD=...the path to libjpl.so , and before that adding the paths to where libjava.so and libjvm.so are located.
Finally got blipkit started from java via jpl
Fri, 2009-10-16 11:17 | by Samuel LampaFinally got blipkit started from java via jpl :)
- (Thanks to Andrew Koster on the SWIPL mailing list, who made me continue researching the LD_PRELOAD trick!)
In order to get the jpl java examples to work, LD_LIBRARY_PATH has to contain:
/usr/lib/jvm/java-1.5.0-sun-1.5.0.19/jre/lib/i386/client/:/usr/lib/jvm/java-1.5.0-sun-1.5.0.19/jre/lib/i386/
Furthermore, CLASSPATH has to be:
/usr/local/lib/pl-5.7.15/lib/jpl.jar:/home/samuel/install/swi-prolog/pl57/packages/jpl/examples/java/Test
Now firing up blipkit works, when calling Java from commandline!!!
On the two main approaches for knowledge handling on the Semantic Web
Wed, 2009-10-07 16:06 | by Samuel LampaEgon pointed me to an interesting blog post on RIF and OWL. The post highlights a situation very relevant for my project: The apparent existence of two main approaches in the field of "knowledge representation and reasoning" (maybe it should be called simply handling?) on the Semantic Web.
Thea - reading OWL2 directly from Prolog
Mon, 2009-10-05 09:56 | by Samuel LampaTalking about BLIPKIT, the Thea library - for reading OWL2 ontologies directly from within SWI-Prolog - seems relevant for this project, as a complement to BLIPKIT. It's written (partly) by the author of BLIPKIT.
Some features I noted:
- SWRL support (SWRL - Semantic Web Rule Language . is a proposed W3C standard for exchange between rule languages on the web)
- Bridge to the java OWL API
- translation of ontologies to Description Logic programs
Switching to Blipkit/BioProlog
Fri, 2009-09-18 14:24 | by Samuel LampaAfter installation problems with DR-PROLOG, and after looking closer to Blipkit/BioProlog (Biomedical Logic Programming Knowledge Integration Kit), after a suggestion by Claes Andersson, I now aim to integrate it instead of DR-PROLOG.